Tuning Guide
Practical guidance for getting the best results from DistributionRegressor.
Step 1: Start with good defaults
For most problems, the following configuration works well out of the box:
model = DistributionRegressor(
n_bins=100,
n_estimators=1000,
learning_rate=0.01,
random_state=42,
)
Step 2: Adjust n_bins for your target range
n_bins controls the resolution of the predicted distribution. Think of it as the
number of "buckets" the model uses to represent the density.
| Scenario | Recommended n_bins |
|---|
| Narrow, well-behaved targets (temperature, standardized scores) |
50 |
| Moderate range (retail sales, housing prices) |
100 |
| Wide range or fine structure (house prices, medical costs) |
150–200 |
| Very wide or heavy-tailed (insurance claims, financial) |
200+ (consider log-transforming the target first) |
Higher n_bins gives finer resolution but increases memory linearly.
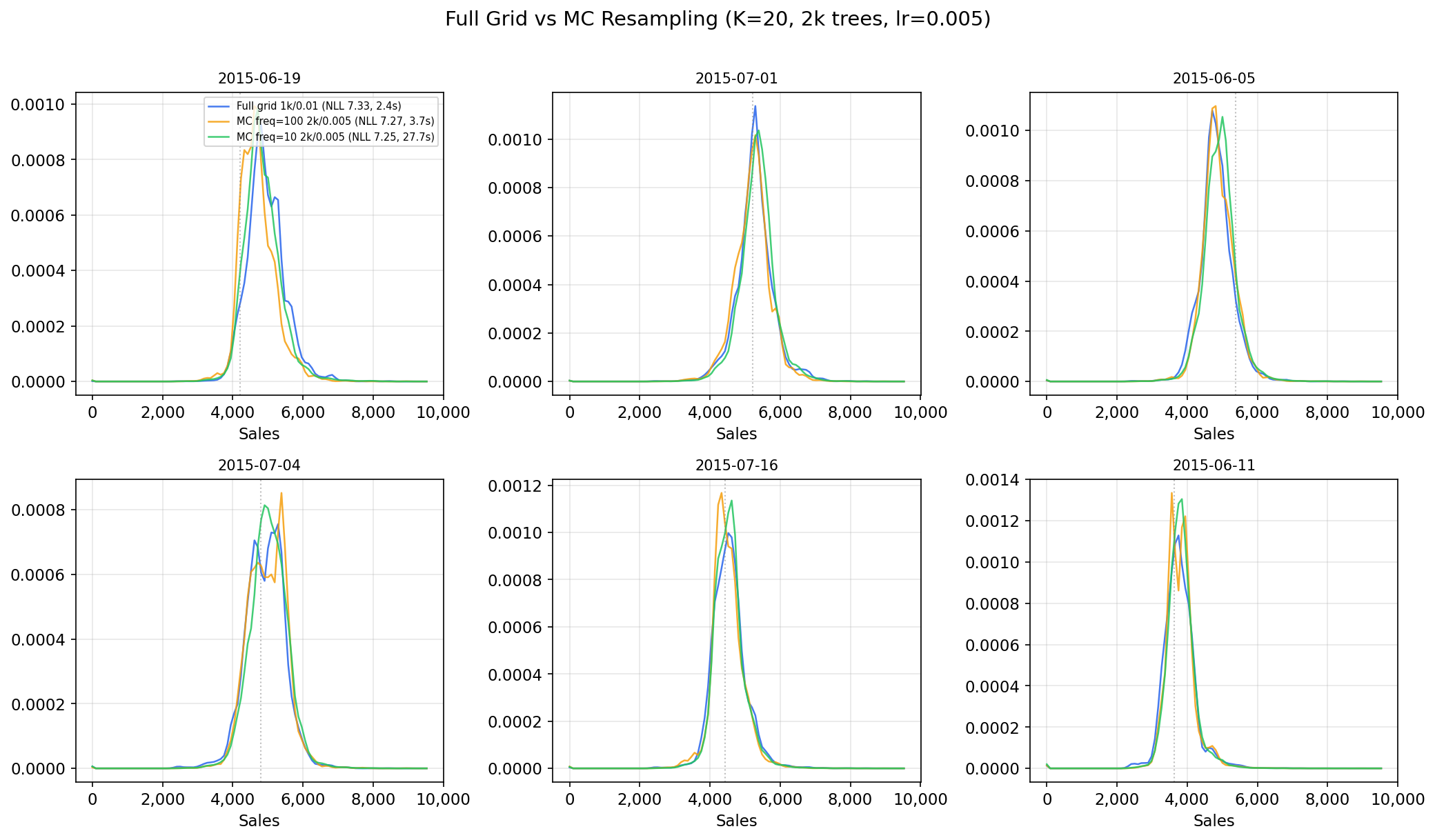
If memory is tight, combine higher n_bins with monte_carlo_training=True
and mc_resample_freq
(see Monte Carlo Training).
Step 3: Tune the LightGBM parameters
Since DistributionRegressor uses LightGBM under the hood, all standard LGBM tuning strategies apply.
Pass any parameter through **kwargs:
model = DistributionRegressor(
n_bins=100,
n_estimators=1000,
learning_rate=0.01,
# Standard LightGBM parameters
num_leaves=63,
max_depth=-1,
subsample=0.8,
colsample_bytree=0.8,
min_child_samples=20,
)
Step 4: Evaluate with proper scoring rules
Use negative log-likelihood (NLL) as your primary evaluation metric for the distribution
quality. Lower NLL means better-calibrated, sharper distributions. Use RMSE or MAE alongside NLL
to evaluate point prediction quality:
# Distribution quality (proper scoring rule)
nll = model.nll(X_test, y_test)
# Point prediction quality
from sklearn.metrics import mean_squared_error
rmse = mean_squared_error(y_test, model.predict(X_test), squared=False)
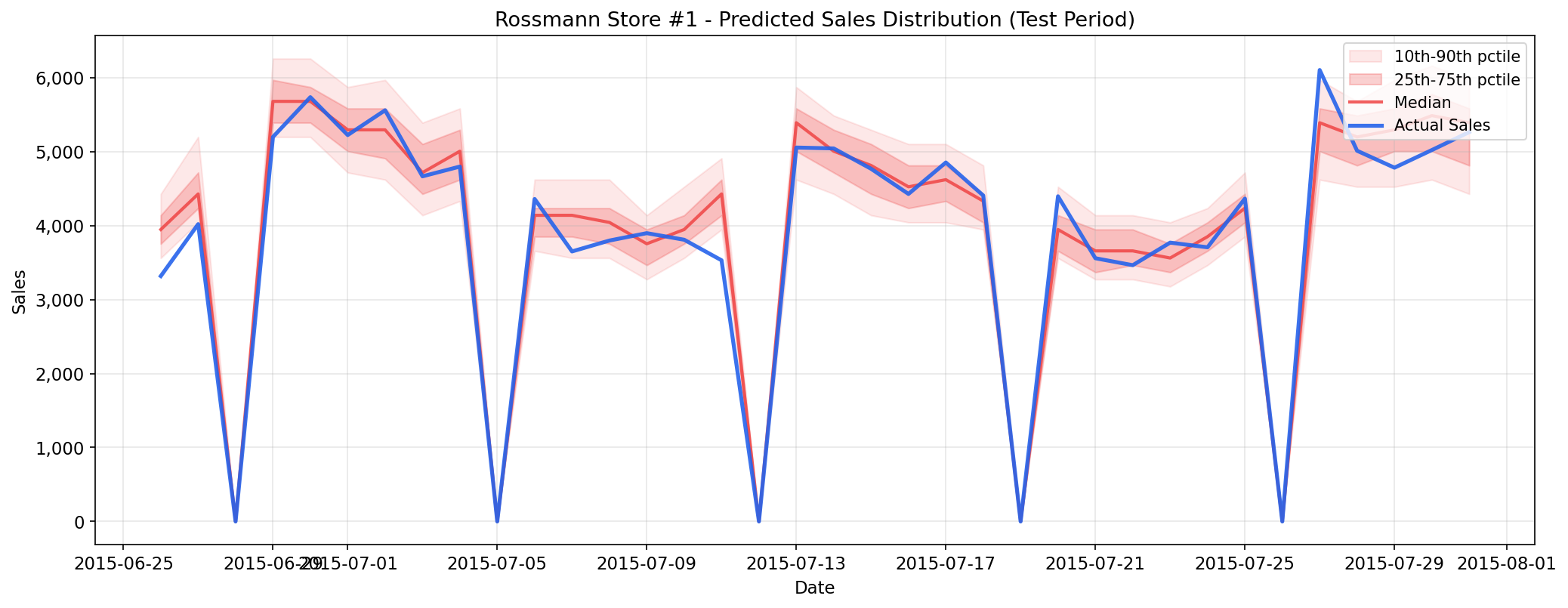
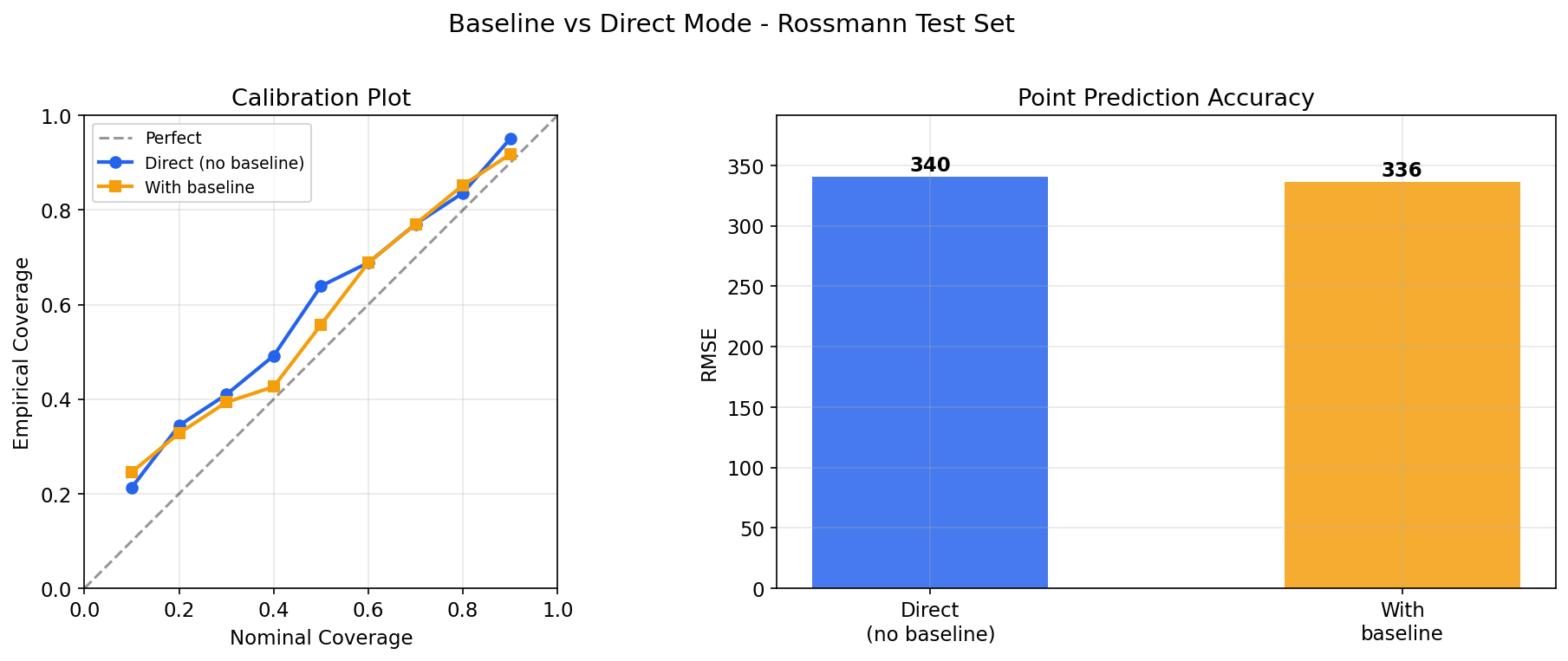
# Calibration check: does 90% interval contain ~90% of true values?
iv = model.predict_interval(X_test, confidence=0.9)
coverage = ((y_test >= iv[:, 0]) & (y_test <= iv[:, 1])).mean()
print(f"90% interval coverage: {coverage:.1%}")
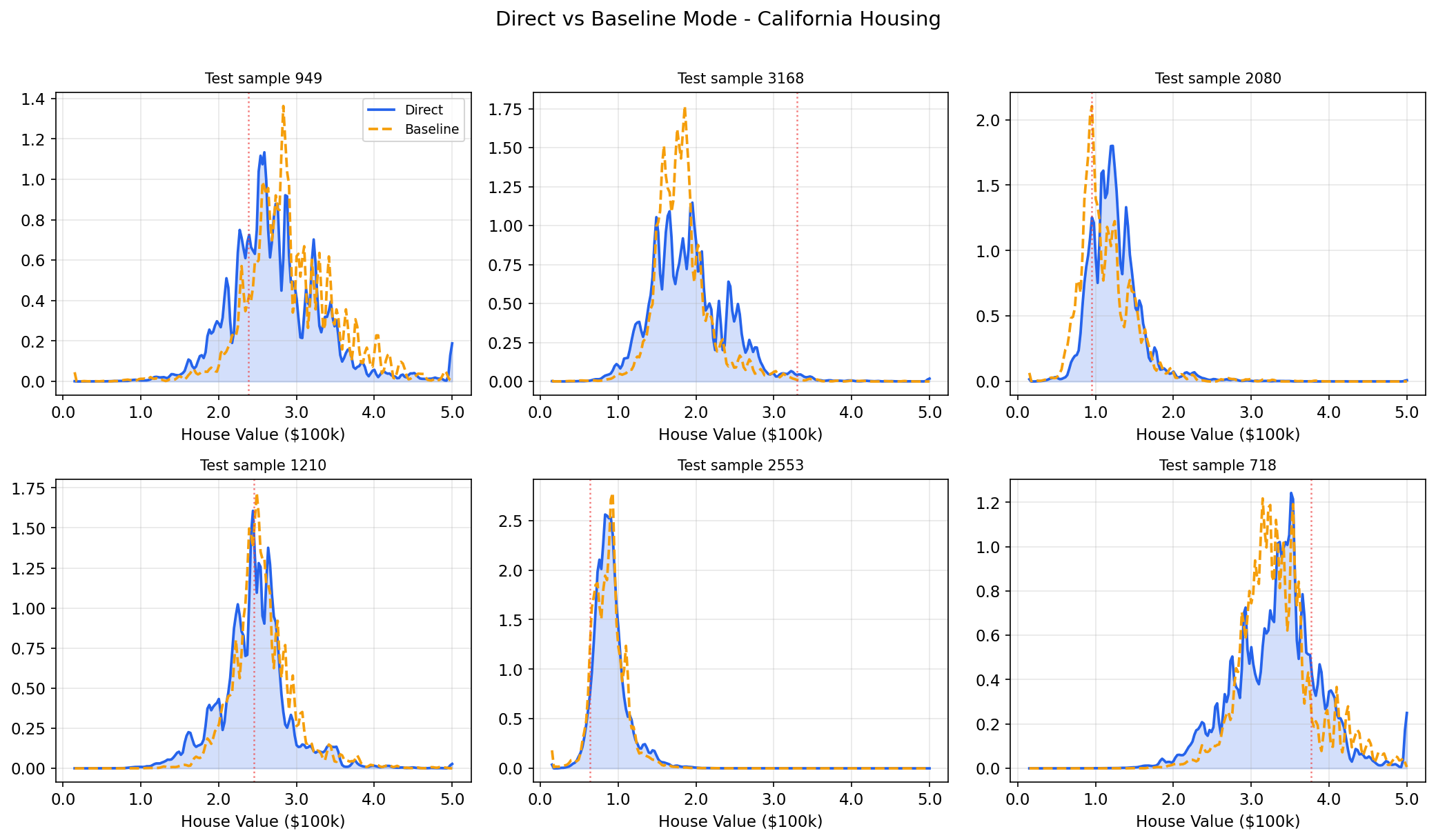
Step 5: Apply output smoothing if needed
If the predicted densities look too jagged (especially at lower n_bins), apply
Gaussian smoothing. You can experiment with this after training without refitting:
# Try different smoothing levels
for sigma in [0, 0.5, 1, 2]:
model.set_output_smoothing(sigma)
nll = model.nll(X_test, y_test)
print(f"sigma={sigma}: NLL={nll:.4f}")
# If target has discrete atoms (e.g., exact zeros), preserve them
model.set_output_smoothing(output_smoothing=1, atom_values=0)
Common pitfalls:
- Too few bins: If
n_bins is much smaller than the effective range of your target,
the predicted distribution will be coarse. Increase n_bins or transform the target.
- Too few estimators: CDF learning benefits from many small steps. If distributions look
underfit (too uniform or too wide), increase
n_estimators or reduce learning_rate.
- Outliers: Extreme outliers stretch the grid range, wasting resolution. Consider
clipping or transforming the target before fitting.